| Doctorant : |
Surbhi DHINGRA
|
| Directeur de thèse : |
Bernard OFFMANN ,
Professeur Université |
| co-directeur de thèse : | CADET Frédéric, Professeur, Université de La Réunion |
| Encadrant : | RAMANATHAN Sowdhamini, NCBS, TIFR (Bangalore, Inde) |
| Financement : |
Région Réunion |
| Date de la soutenance : |
vendredi 12 juin 2020, 13h00 |
| Modalité : |
|
| Jury : |
- Président de jury : NARAYASWAMY Srinivasan, Professeur, Indian Institute of Science (Bangalore, Inde)
- Rapporteur : CAMPROUX Anne-Claude, Professeure des universités, Université de Paris
- Rapporteur : MARTIN Juliette, CR CNRS HDR, Université de Lyon
- Directeur de thèse :
Bernard OFFMANN ,
Professeur Université
- co-directeur de thèse : CADET Frédéric, Professeur, Université de La Réunion
- Encadrant : RAMANATHAN Sowdhamini, NCBS, TIFR (Bangalore, Inde)
|
La modélisation des structures protéiques en l’absence d’homologie est appelée modélisation sans matrice, modélisation libre ou modélisation ab initio. À ce jour, ce type de modélisation reste le plus grand défi du domaine.
Durant cette thèse, nous avons apporté une contribution méthodologique pour relever ce défi en explorant l’utilisation d’un alphabet structurel connu sous le nom de blocs protéiques (BP). Il est maintenant possible de prédire le squelette carbonné d’une protéine sous forme de séquence de BPs de n’importe quelle protéine en utilisant des outils comme PB-kPRED. Notre travail a consisté à trouver une méthode pour passer de la séquence de BPs prédite à la structure 3D.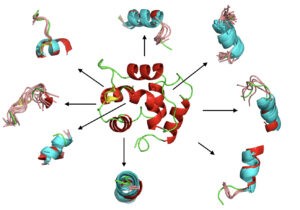
Nous avons donc développé une stratégie qui part d’une requête sous forme de séquences de BP prédites et recherche dans une base de données de structures non homologues des fragments de BPs similaires.
Notre stratégie tente ensuite d’assembler des fragments sélectionnés pour générer des modèles 3D. Pour cela, nous avons tenté d’exploiter les propriétés inhérentes de Modeller en la contraignant à utiliser des fragments de BPs et à utiliser des prédictions de structures secondaires et de contact comme contraintes externes supplémentaires.
Nous avons validé les résultats de cette stratégie sur un sous-ensemble de petites protéines cibles issues de CASP13 sur la base des scores couramment utilisés (GDT_TS et TM-score) pour les protocoles de modélisation gratuits. Nous avons pu montrer que des modèles de bonne qualité peuvent être générés pour de petites protéines en utilisant une combinaison de fragments à base de PB et de Modeller. Ce travail ouvre de nouvelles perspectives d’application des blocs protéiques pour solutionner l’épineux problème du repliement des protéines.
Publications
2020
Dhingra, Surbhi; Sowdhamini, Ramanathan; Sanejouand, Yves-Henri; Cadet, Frédéric; Offmann, Bernard
Customised fragment libraries for ab initio protein structure prediction using a structural alphabet Article de journal
Dans: arXiv:2005.01696, 2020.
@article{Dhingra2020,
title = {Customised fragment libraries for ab initio protein structure prediction using a structural alphabet},
author = {Surbhi Dhingra and Ramanathan Sowdhamini and Yves-Henri Sanejouand and Frédéric Cadet and Bernard Offmann},
url = {https://arxiv.org/pdf/2005.01696.pdf},
year = {2020},
date = {2020-05-01},
journal = {arXiv:2005.01696},
abstract = {Motivation: Computational protein structure prediction has taken over the structural community in past few decades, mostly focusing on the development of Template-Free modelling (TFM) or ab initio modelling protocols. Fragment-based assembly (FBA), falls under this category and is by far the most popular approach to solve the spatial arrangements of proteins. FBA approaches usually rely on sequence based profile comparison to generate fragments from a representative structural database. Here we report the use of Protein Blocks (PBs), a structural alphabet (SA) to perform such sequence comparison and to build customised fragment libraries for TFM. Results: We demonstrate that predicted PB sequences for a query protein can be used to search for high quality fragments that overall cover above 90% of the query. The fragments generated are of minimum length of 11 residues, and fragments that cover more than 30% of the query length were often obtained. Our work shows that PBs can serve as a good way to extract structurally similar fragments from a database of representatives of non-homologous structures and of the proteins that contain less ordered regions.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Motivation: Computational protein structure prediction has taken over the structural community in past few decades, mostly focusing on the development of Template-Free modelling (TFM) or ab initio modelling protocols. Fragment-based assembly (FBA), falls under this category and is by far the most popular approach to solve the spatial arrangements of proteins. FBA approaches usually rely on sequence based profile comparison to generate fragments from a representative structural database. Here we report the use of Protein Blocks (PBs), a structural alphabet (SA) to perform such sequence comparison and to build customised fragment libraries for TFM. Results: We demonstrate that predicted PB sequences for a query protein can be used to search for high quality fragments that overall cover above 90% of the query. The fragments generated are of minimum length of 11 residues, and fragments that cover more than 30% of the query length were often obtained. Our work shows that PBs can serve as a good way to extract structurally similar fragments from a database of representatives of non-homologous structures and of the proteins that contain less ordered regions.
Dhingra, Surbhi; Sowdhamini, Ramanathan; Cadet, Frédéric; Offmann, Bernard
A glance into the evolution of template-free protein structure prediction methodologies Article de journal
Dans: Biochimie, vol. 175, p. 85 - 92, 2020, ISSN: 0300-9084.
@article{DHINGRA202085,
title = {A glance into the evolution of template-free protein structure prediction methodologies},
author = {Surbhi Dhingra and Ramanathan Sowdhamini and Frédéric Cadet and Bernard Offmann},
url = {http://www.sciencedirect.com/science/article/pii/S0300908420300961},
doi = {https://doi.org/10.1016/j.biochi.2020.04.026},
issn = {0300-9084},
year = {2020},
date = {2020-01-01},
journal = {Biochimie},
volume = {175},
pages = {85 - 92},
abstract = {Prediction of protein structures using computational approaches has been explored for over two decades, paving a way for more focused research and development of algorithms in comparative modelling, ab intio modelling and structure refinement protocols. A tremendous success has been witnessed in template-based modelling protocols, whereas strategies that involve template-free modelling still lag behind, specifically for larger proteins (>150 a.a.). Various improvements have been observed in ab initio protein structure prediction methodologies overtime, with recent ones attributed to the usage of deep learning approaches to construct protein backbone structure from its amino acid sequence. This review highlights the major strategies undertaken for template-free modelling of protein structures while discussing few tools developed under each strategy. It will also briefly comment on the progress observed in the field of ab initio modelling of proteins over the course of time as seen through the evolution of CASP platform.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Prediction of protein structures using computational approaches has been explored for over two decades, paving a way for more focused research and development of algorithms in comparative modelling, ab intio modelling and structure refinement protocols. A tremendous success has been witnessed in template-based modelling protocols, whereas strategies that involve template-free modelling still lag behind, specifically for larger proteins (>150 a.a.). Various improvements have been observed in ab initio protein structure prediction methodologies overtime, with recent ones attributed to the usage of deep learning approaches to construct protein backbone structure from its amino acid sequence. This review highlights the major strategies undertaken for template-free modelling of protein structures while discussing few tools developed under each strategy. It will also briefly comment on the progress observed in the field of ab initio modelling of proteins over the course of time as seen through the evolution of CASP platform.
Lien


